Microbiome Research
HICAHS experts apply microbiome research to livestock production settings with the goal of improving worker health.
In addition to collaborating on research projects funded though the Center’s CDC/NIOSH funded research core, HICAHS researchers also work on projects related to agricultural worker health and safety supported by other funding mechanisms. One such focus area has been the microbiome’s role in occupational health: specifically, characterizing the microbes found in livestock workers and livestock production environments and evaluating their impact on health.
Microbes are present in all corners of the world in staggering numbers. For example, the ocean floor is home to 1029 microbial cells, or approximately 0.6% of the Earth’s total living biomass. The human body harbors an astounding number of microbes; the gastrointestinal tract alone is colonized by trillions of interacting microbes. Furthermore, microbes are highly concentrated in the near-surface atmosphere—millions of cells may be present in one cubic meter of air.
Microbes are essential and intimately linked to fundamental processes that promote human, animal and environmental health. Conversely, there are subsets of microbes that are known pathogens. Additionally, nonviable microbial particulate matter may also trigger allergic and inflammatory responses. Community alterations and environmental perturbations may facilitate and promote the presence and growth of these harmful species.
Aerobiome Discovery Network
HICAHS Director, Stephen Reynolds, and HICAHS Collaborators, Sheryl Magzamen and Joshua Schaeffer, participate in the Aerobiome Discovery Network. This research team works to converge previously unlinked expertise in atmospheric science, infectious diseases/pathogens of plants, animals, people, and ecology, epidemiology and microbiome/genomic sciences at Colorado State University to explore the fascinating field of aerobiology – the study of organic particles which are passively transported by the air. The Aerobiome Discovery Network is funded through the School of Global Environmental Sustainability at Colorado State University.
Antimicrobial resistant bacteria: Exposure and Health of Cattle Workers
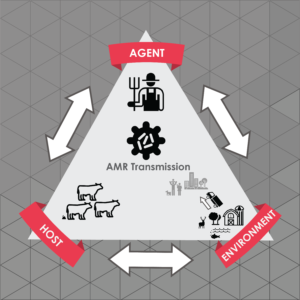 Livestock workers are at the frontline of exposure to agriculture-generated bioaerosols containing a diverse mixture of respiratory pathogens. The misuse and overuse of antibiotics in agriculture has been posited as a potential driver in the accelerated development of antibiotic resistant genes (ARG) that may be present in these bioaerosols. We hypothesize that antibiotic-resistant bacteria are evolving at the human-animal interface, which has broad health and economic implications for agricultural producers. We will address critical questions that remain about the impacts of worker exposure to respiratory pathogens and transmission to animals, farms and the larger community. Specifically, we will ascertain if agricultural workers act as vectors to introduce ARG and viruses in the community and farm. The research objective is to ascertain if ARG and viruses are underlying risk factors for subclinical markers of respiratory inflammation. The central hypothesis is that the nasal microbiome of cattle workers will have distinct microbial communities and genetic constitution that are associated with measurable levels of inflammation and airway resistance as compared to occupational controls and housemates. Full project details.
Livestock workers are at the frontline of exposure to agriculture-generated bioaerosols containing a diverse mixture of respiratory pathogens. The misuse and overuse of antibiotics in agriculture has been posited as a potential driver in the accelerated development of antibiotic resistant genes (ARG) that may be present in these bioaerosols. We hypothesize that antibiotic-resistant bacteria are evolving at the human-animal interface, which has broad health and economic implications for agricultural producers. We will address critical questions that remain about the impacts of worker exposure to respiratory pathogens and transmission to animals, farms and the larger community. Specifically, we will ascertain if agricultural workers act as vectors to introduce ARG and viruses in the community and farm. The research objective is to ascertain if ARG and viruses are underlying risk factors for subclinical markers of respiratory inflammation. The central hypothesis is that the nasal microbiome of cattle workers will have distinct microbial communities and genetic constitution that are associated with measurable levels of inflammation and airway resistance as compared to occupational controls and housemates. Full project details.
Funding: NIOSH R01, 2020-2024
Contacts: Joshua Schaeffer, Sheryl Magzamen, Stephen Reynolds
CHROME
The Collaborative Health Research on the Microbiome and the Environment, or CHROME, focuses on the synergistic relationships between human, animal and environmental health. With the advent of next-generation sequencing, we are shifting the traditional paradigm of microbiology and industrial hygiene to high-throughput metagenomic approaches to better characterize and understand microbial niches, co-occurrences and competitive exclusions in the occupational and environmental setting. With the capabilities of bioinformatics and data pipelines, analytic approaches can move beyond basic descriptive statistics to better anticipate and develop interventions that effectively reduce or prevent occupational exposures and environmental stressors.
- Bioaerosol Exposures and Models of Human Responses in Dairies and Cattle Feedlots
Project PI: Stephen Reynolds - Modeling and Predicting Microbiomes in Dairies: A Metagenomic Assessment of Bioaerosols
Project PI: Joshua Schaeffer - Exposure Assessment of Bioaerosols in an Australian Dairy
Project PI: Sue Reed - E. coli O157 Shedding and Antimicrobial Susceptibility on Colorado Dairies
Project PI: Craig McConnel - Characterizing Biological Pollutants in Agricultural Runoff at Colorado Dairies
Project PI: Sheryl Magzamen